Our technology
C-FISH stands for Computerized- Fluorescence In Situ Hybridization.
The basis for C-FISH is FISH. A technology that was first developed late last century.
FISH, or Fluorescence In Situ Hybridization, is a powerful molecular biology technique used for the detection and visualization of specific nucleic acid sequences within cells, tissues, or microorganisms. It is particularly valuable in identifying and characterizing microorganisms, such as bacteria, viruses, and fungi, in various biological samples.
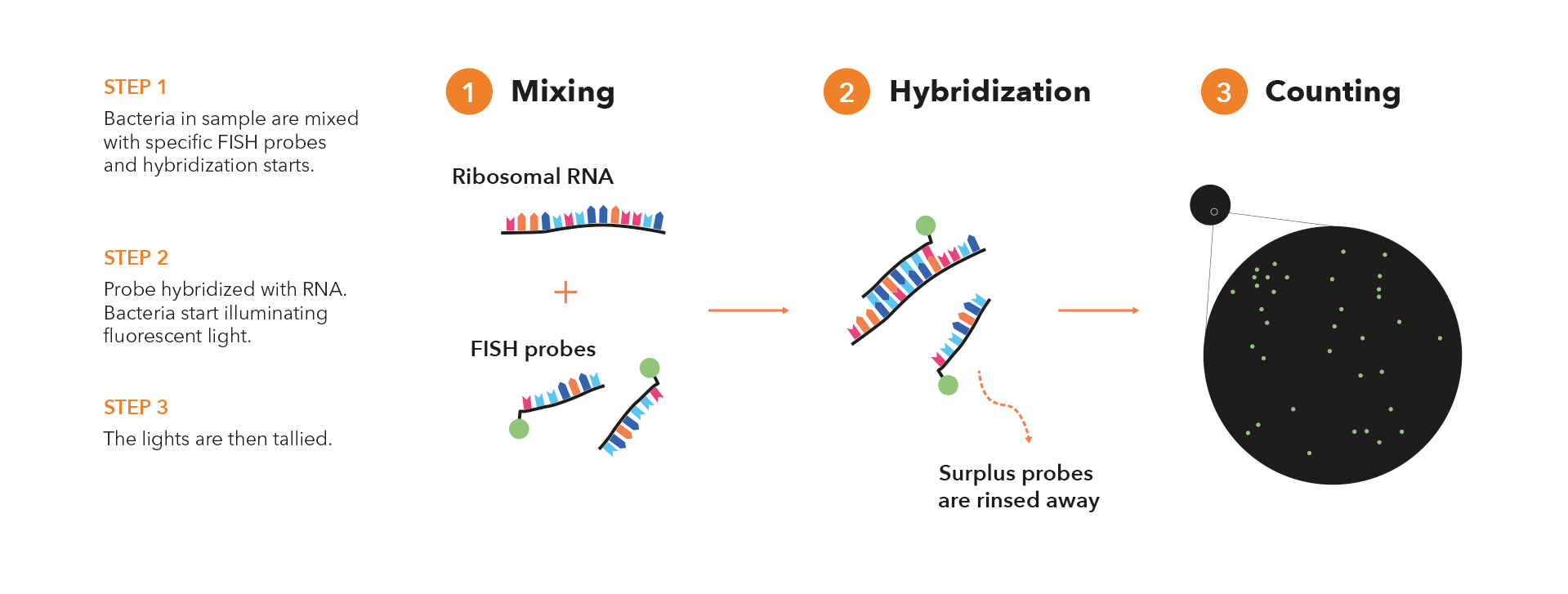
Here’s how FISH works in a few basic steps:
- Probe Design: In FISH, a specific nucleic acid probe is designed to target a particular genetic sequence of interest. This probe is usually a short single-stranded DNA or RNA molecule that is complementary to the target sequence. The probe is labeled with a fluorescent molecule, which allows it to emit light when exposed to specific wavelengths of light.
- Sample Preparation: The sample containing the microorganisms of interest is typically fixed using a fixative agent. This could be a clinical specimen, or an environmental sample.
- Hybridization: The labeled probe is then applied to the sample and allowed to hybridize (bind) with its complementary target sequence within the microorganisms’ genetic material. The hybridization process requires specific temperature and buffer conditions to ensure proper binding.
- Washing and Detection: After hybridization, the sample is washed to remove any unbound probes. The remaining bound probes indicate the presence and location of the target microorganisms. The labeled probes emit fluorescence when excited by specific wavelengths of light, allowing the microorganisms to be visualized under a fluorescence microscope.
- Microscopy and Imaging: The sample is examined using a fluorescence microscope equipped with appropriate filters to capture the emitted fluorescence. Each microorganism or cell containing the target sequence will emit fluorescence, and their presence and distribution can be observed and analyzed.
FISH in general has been widely used in various fields, including clinical microbiology, environmental science, and research. It helps researchers and clinicians gain a deeper understanding of microbial communities, pathogenic infections, and microbial ecology.
C-FISH has several advantages for microorganism detection:
Specificity: C-FISH offers high specificity because it relies on the complementary base pairing between the probe and the target sequence. This ensures that only the intended microorganisms are detected.
Sensitivity: C-FISH is able to detect the target on a single cell level due the automated UHD digital imaging, thus ensuring the best possible sensitivity for microorganisms.
Visualization: C-FISH allows visualization of the spatial distribution of microorganisms within a sample, providing insights into their localization and interactions.
Cultivation-Independent: C-FISH can detect microorganisms without the need for prior cultivation or growth, which is especially beneficial for studying organisms that are difficult to culture.
Rapid Results: C-FISH provides relatively quick results compared to traditional culture-based methods, making it valuable for clinical diagnostics and environmental monitoring.
Microbial Activity: C-FISH is able to determine microbial activity, a new ability in the field of microorganism detection.
Fluorescence in situ hybridization (FISH) is a powerful molecular biology technique that allows us to visualize and map the location of specific DNA or RNA sequences within cells and tissues. It has greatly contributed to our understanding of genetics, gene expression, chromosomal abnormalities, and various biological processes. The history of FISH is marked by key developments and milestones:
- Early Concepts and Developments (1960s-1970s):
- The groundwork for FISH was laid in the 1960s when researchers began experimenting with nucleic acid hybridization techniques. Hybridization involves pairing complementary DNA or RNA strands together.
- In 1969, John Cairns used radioactive probes to detect DNA sequences in the bacterial genome, marking one of the earliest instances of in situ hybridization.
- In the early 1970s, researchers like Mary-Lou Pardue and Joseph Gall used radioactive RNA probes to visualize specific gene sequences on Drosophila (fruit fly) chromosomes.
- Introduction of Non-Radioactive Probes (1980s):
- The use of radioactive probes posed safety and disposal concerns, which led to the development of non-radioactive probes for FISH.
- In 1980, Louise Brown and John Smith published a method using biotin-labeled probes for in situ hybridization.
- Throughout the 1980s, researchers explored alternative labeling techniques, such as digoxigenin-labeled probes, that produced detectable signals without using radioactivity.
- Fluorescence Labeling and Confocal Microscopy (1980s-1990s):
- The breakthrough came with the incorporation of fluorescent dyes as labels for FISH probes. This allowed for easier detection and visualization of hybridization events.
- In 1986, John W. Sedat and David A. Agard introduced wide-field epifluorescence microscopy to image FISH probes.
- In 1989, Robert Singer and colleagues developed the method of fluorescence in situ hybridization using directly labeled DNA probes.
- Chromosome Painting and Spectral Karyotyping (SKY) (1990s-2000s):
- Researchers began using multiple fluorescent probes to simultaneously label different chromosomes, a technique known as chromosome painting.
- In 1995, the Spectral Karyotyping (SKY) technique was introduced, allowing for the simultaneous visualization of all chromosomes using different fluorescent colors.
- These techniques revolutionized the study of chromosomal abnormalities and genetic variations.
- Advancements in Probe Design and Automation (2000s-Present):
- The Biotrack platform and C-FISH technology is an automated FISH system and combines high-throughput techniques allowing to analyze large numbers of samples more efficiently with UHD imaging and AI driven algorithms.
- Biotrack is working on (ongoing) advancements in probe design, labeling technologies, digital imaging and AI driven algorithms to further improve the unprecedented sensitivity, specificity, and speed of FISH.
- FISH has found applications in cancer diagnostics, prenatal testing, microbial ecology, and various fields of research.
The history of FISH illustrates the evolution of molecular techniques and their profound impact on our understanding of genetics and biology. FISH continues to be a fundamental tool for studying genome organization, gene expression, and chromosomal abnormalities in various organisms and cell types.